Solutions
Affinity Screen
The Orbit Discovery Affinity Screening services offers the opportunity to apply a proprietary technology platform that can interrogate populations of binding peptides using FACS. This capability offers a significant throughput benefit beyond more traditional screening techniques, such as phage display or ribosomal display. As well as screening against soluble targets, the platform can be combined with other internal capabilities to offer a unique ability to address cell surface targets in a primary peptide screen, a particularly useful benefit for identifying targeting peptides.
The bead-based technology enables the construction of massive libraries with peptides displayed on their surface. Each bead carries a unique DNA sequence, which is translated into peptide using in vitro transcription translation (IVTT) performed in an emulsion (emIVTT). Each droplet of the emulsion acts as a micro-compartment which allows translation of the DNA on each individual bead to occur separately. The result is a library of unique beads each displaying thousands of copies of peptide, providing massive chemical diversity and the ability to perform rapid screens for high affinity binders against soluble or more complex target proteins, such as GPCRs and ion channels.
The platform is flexible, allows for different library structures (linear, cyclic, helical), peptide lengths, diversity, and chemistries. The high diversity that can be covered produces an abundance of hits that can be further optimised as therapeutic candidates. Binders in the nanomolar range can be detected direct from the screen, weak binders in the micromolar range can also be identified. Binders are analysed for conserved motifs and structures, these can be synthesised, and binding confirmed (peptide microarray, thermal shift, SPR or cell assays can be performed) prior to taking into further medicinal chemistry. Alternatively, binders can be used to design smaller more refined libraries that can be taken into cell based functional screening.
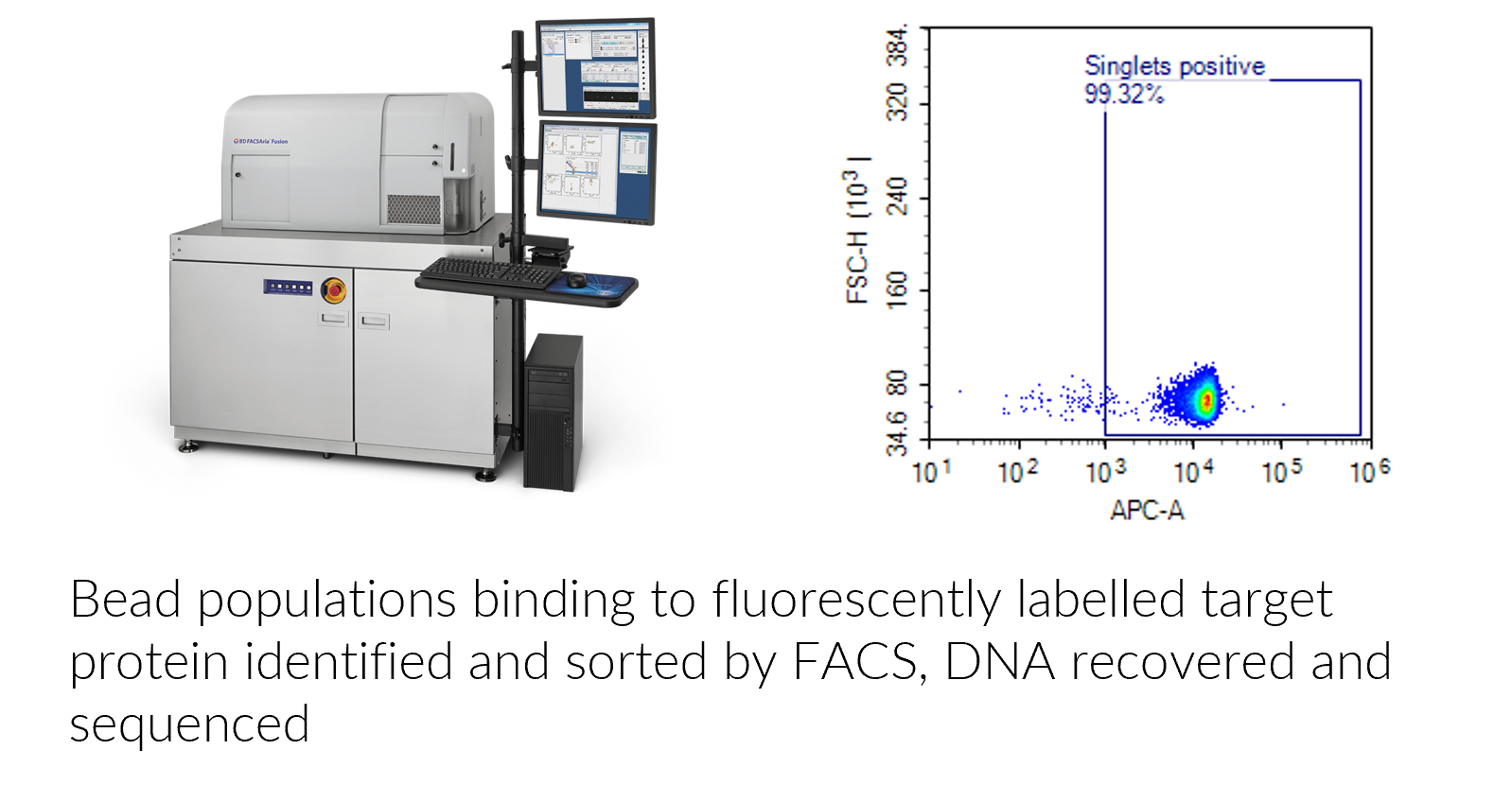
DNA encoded peptide MDM2 library screened by FACS for target binders
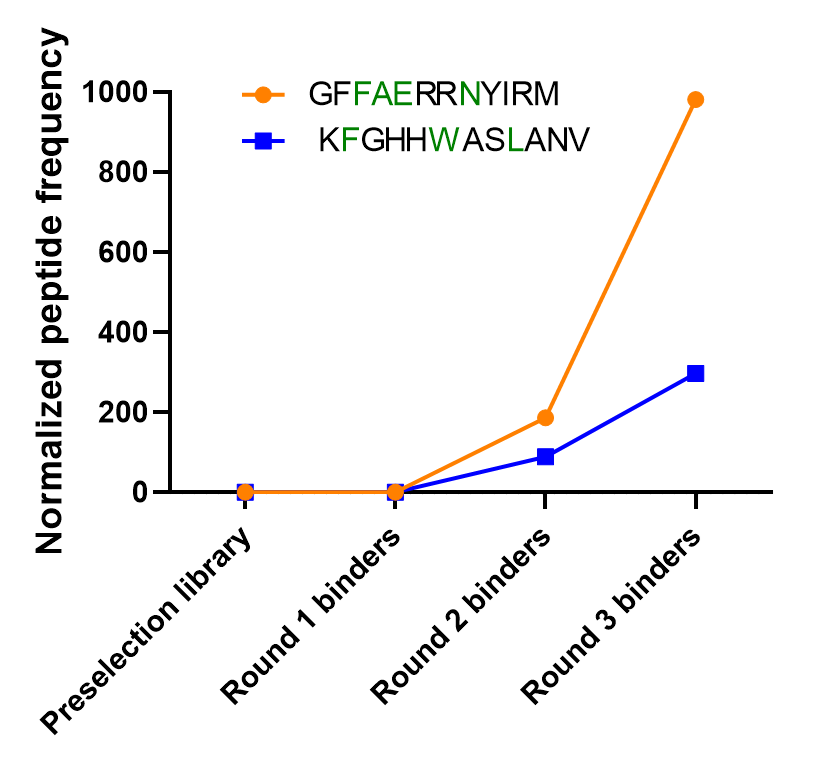
MDM2 binding peptides enriched through screening rounds
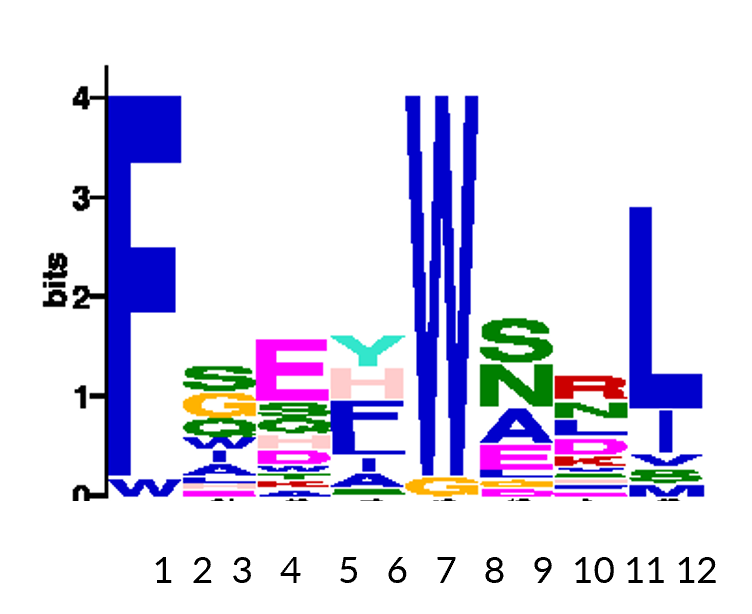
Bionformatic analysis of hits reveals important motifs
Key benefits of Orbit's affinity screening
-

Sensitivity ~ nM to uM binders
-

Highly diverse massive libraries
-

Cell based screening
For Affinity Screening services contact us
Find out how Orbits’s technology solution can support your business.